Highlights
Researchers reveal novel catalytic mechanism of Nonribosomal peptide synthetases
Nonribosomal peptide synthetases (NRPSs) are mega-enzymes that can assemble proteinogenic and nonproteinogenic amino acids into a vast variety of peptides with antibacterial, antiviral, immunosuppressor, or antitumor activities. NRPSs synthesize peptides in a modular and pipelined fashion, and each module adds one amino acid to the nascent peptide. A basic NRPS module consists of a condensation domain, an adenylation domain, and a peptidyl carrier protein domain. While the role of each single domain in catalytic activity is well accepted, whether the condensation (C) domain and the adenylation (A) domain cooperate with each other for the efficient catalytic activity remains poorly understood.
In a study published in Structure, the research group led by Prof. ZHANG Cheng-Cai from the Institute of Hydrobiology (IHB) of the Chinese Academy of Sciences reported the molecular mechanism of the C domain in helping the A domain of the same NRPS module to select specific substrate efficiently during catalysis.
Microcystins (MCs) are typical nonribosomal peptides produced by a diverse range of bloom-forming freshwater cyanobacteria. They are the most common hepatotoxins present in aquatic environment. Through biochemistry studies, Zhang's team first identified that the presence of the C domain enhances the activity of the cognate A domain in different MC NRPS modules.
To uncover the underlying catalytic mechanisms, the researchers solved two high-resolution X-ray structures of the MC NRPS modules representing two newly identified conformations in the NRPS catalytic cycle. Crystal structural analysis, mutagenesis of key residues and cysteine-based crosslinking studies demonstrated that the dynamic interaction between the C and the A domains in these modules are mediated by the conserved “RXGR” motif, and this interaction is important for the adenylation activity.
“Most importantly, the “RXGR” motif-mediated dynamic interaction and its functional regulation found in this study is prevalent in different NRPSs modules. This provides new insights into the catalytic mechanism of NRPSs and their engineering strategy for synthetic peptides with new structures and properties.” Prof. Zhang said.
Based on the findings, Zhang’s team propose a model on the control mechanism of the adenylation activity in a NRPS module mediated by dynamic interaction between the two domains. When the substrate of a NRPS module binds at the pocket of the Acore domain, it triggers a change at a right angle of the “hinge” residues located between the two subdomains. Subsequently, the dynamic interaction between the “RXGR” motif and the C domain guides the rotational direction of the Asub domain, enabling a more efficient state transition and a high adenylation efficiency of the A domain. When the dynamic interaction between the “RXGR” motif and the C domain was disrupted, the Asub domain rotates in a random fashion, thereby resulting in a low adenylation efficiency of the A domain in a NRPS module.
This work was funded by National Key R&D Program of China (project number 2018YFA0903100) and the Youth Innovation Promotion Association CAS.
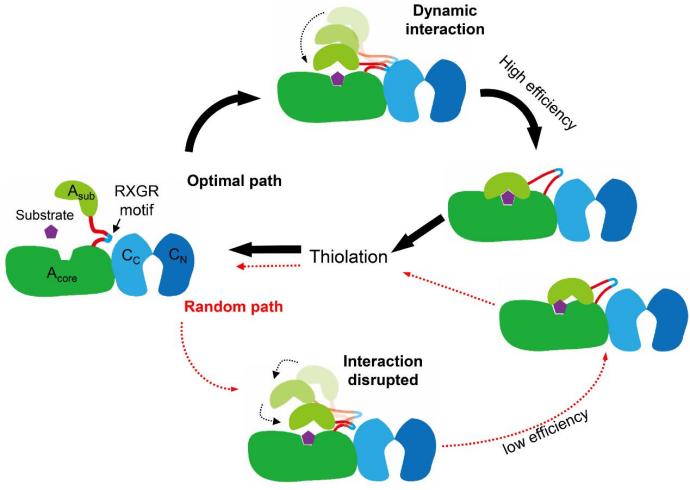
A model for the control mechanism of the adenylation activity in a NRPS module, mediated by dynamic interaction between the A and C domains (Image by IHB)
(Editor. WANG Hongxia)