
Newsroom
Scientists Discover First Functional Divergence of Diacylglycerol Acyltransferases in Microalga
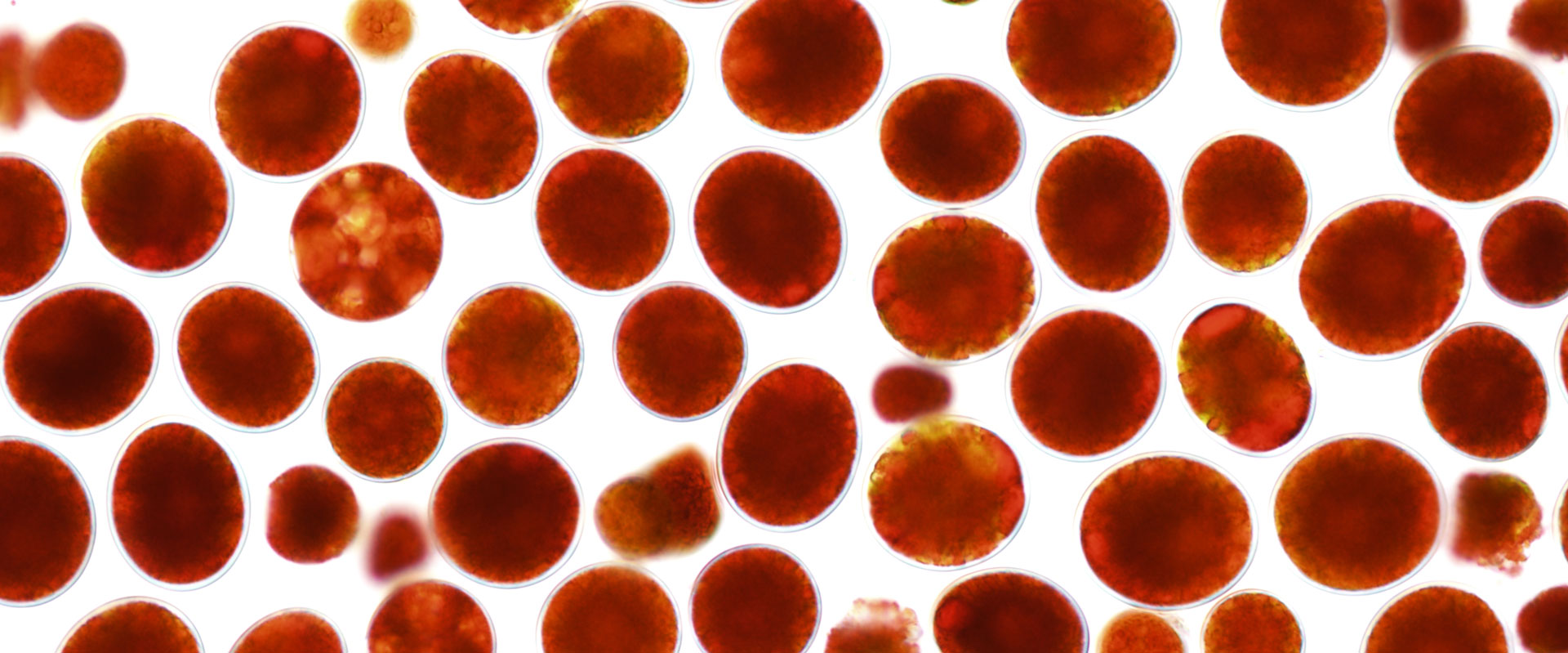
Red cyst of Haematococcus pluvialis (credit: IHB)
Researchers from the Institute of Hydrobiology (IHB) of Chinese Academy of Sciences recently revealed, for the first time, that microalgal diacylglycerol acyltransferases (DGAT) can function as a lysophosphatidic acyltransferase (LPAAT) both in vitro and in vivo while losing its original function as DGAT.
The research findings were published in the Journal of Experimental Botany, providing essential clues for understanding the functional divergence of DGAT family in microalga.
The research team identified and cloned from the green microalga Haematococcus pluvialis five DGAT-encoding genes. They found that four encoded HpDGATs showed triacylglycerol synthase activities in yeast functional complementation analyses, with an exception for one of the type II DGAT encoding genes, HpDGTT2.
They also purified the hydrophobic recombinant HpDGTT2 protein in soluble form and found that it functions as a LPAAT via enzymatic assay. This was the first time for scientists to perform function characterization with a soluble DGAT in an in vitro assay.
Additionally, researchers found that when this gene was introduced into the green microalga Chlamydomonas reinhardtii, the cellular growth was retarded with enlarged cell size and enhanced triacylglycerol accumulation, which were identical to the phenotypes of transgenic strains overexpressing CrLPAAT.
In-depth sequence alignment and topology analysis indicated that the existence and particular position of the third transmembrane domain in HpDGTT2 dramatically changed its topological structure, which may be attributed to the functional switch of HpDGTT2 from DGAT to LPAAT that involves a similar topology.
DGAT catalyzes the final and key step of Triacylglycerol (TAG) biosynthesis, and plays crucial roles in several biological processes such as development of seeds and pollen, metabolism of leaves. In animals, DGAT is considered to be the medicinal target for curing obesity and diabetes, and is also the key target gene for enhancing the lipid production in microalga.
In eukaryotic organisms, DGAT are mainly encoded by two distinct gene families, referred to as DGAT1 and DGAT2. The DGAT2 gene family was found to be extensively expanded in microalgal genomes. In the model green alga Chlamydomonas reinhardtii, there are five copies of the DGAT2 gene, while the heterokont microalga Nannochloropsis oceanica possesses 11 DGAT2 genes.
Moreover, many algal DGAT2 isoforms could not complement the TAG-deficient phenotype of Saccharomyces cerevisiae H1246, a tool frequently used to verify the function of DGAT, nor could they exhibit any enzymatic activity in vitro. These findings raise a crucial question about the authentic functions of these cryptic DGAT2 genes in microalgae.
This above study conducted by IHB thus provides a framework for dissecting uncharacterized DGATs, and could pave the way to decrypting the structure–function relationship of this large group of enzymes that are critical to lipid biosynthesis.