
Newsroom
Researchers Reveal Carbon Cycle in Microbial Ecosystems of Biological Soil Crusts
As greenhouse effects become increasingly prominent, soil carbon has been a major focus of research on climate change. Soil microorganisms are the key groups that drive soil carbon transformation. Due to the complexity of factors such as microbial physiology, the composition of organic compounds in soils, and variation among redox forms, the pattern and process of soil carbon cycle at the microbial community level can be difficult to study using metagenomics.
For studies of soil ecosystems using metagenomics, biocrusts are ideal for modeling because they are widely distributed in arid regions, account for > 40% of the global land surface, and are dominated by cryptophytes like Cyanobacteria, lichens, and mosses. They can also prevent wind erosion of sandy soils, thus promoting ecosystem development and succession.
In a study published online in Soil Biology and Biochemistry, a research team led by Prof. HU Chunxiang from the Institute of Hydrobiology (IHB) of the Chinese Academy of Sciences explored the community-based carbon cycle overall patterns and successional variation in biocrusts, and the relationships among C-cycle processes, environmental variables, and microorganismal groups.
To achieve this, the researchers used four types of biocrusts that represent different successional stages, i.e., cyanobacteria-dominated crusts, A; cyanolichen-dominated crusts, C; chlorolichen-dominated crusts, G; and moss-dominated crusts, M. They were collected repeatedly from Shapotou District in the Tengger Desert over the past four years.
The researchers first observed a low abundance of genes related to light-driven inorganic carbon fixation, and high abundance of genes related to the chemical energy-driven degradation of macromolecular organic carbon (OC), fermentation, aerobic respiration, and CO oxidation using metagenomic sequencing.
“For OC decomposition, we found that genes mediating starch/glycogen and cellulose degradation were most abundant during the initial complex OC degradation, as were genes mediating fermentation during terminal steps of OC decomposition,” said Prof. Hu.
To assess successional changes in carbon cycle, the researchers further combined the metagenomic data with absolute quantification via GeoChip and key enzyme activity measurements. “We observed that inorganic carbon fixation, fermentation, CH4 oxidation, and both starch/glycogen and peptidoglycan degradation decreased during succession, whereas several high efficiency processes, as well as CO oxidation and most types of OC degradation, increased,” said Prof. Hu.
In addition, using co-occurrence networks, the researchers revealed that C-cycle in biocrusts comprises an assimilation module, akin to primary production, and a dissimilation module, comparable to secondary production. They also found that dynamic changes in the relationships between C-cycle pathways and microbial community composition occurred during succession. “The two C-cycle modules were connected by the Calvin-Benson-Bassham cycle, ethanol and propionate fermentation, and they were balanced by drought and salinity,” Prof. Hu explained.
This study improved the understanding of C-cycle pathways and regulatory mechanisms in biocrust succession, and laid a groundwork for future multi-omics studies of these systems.
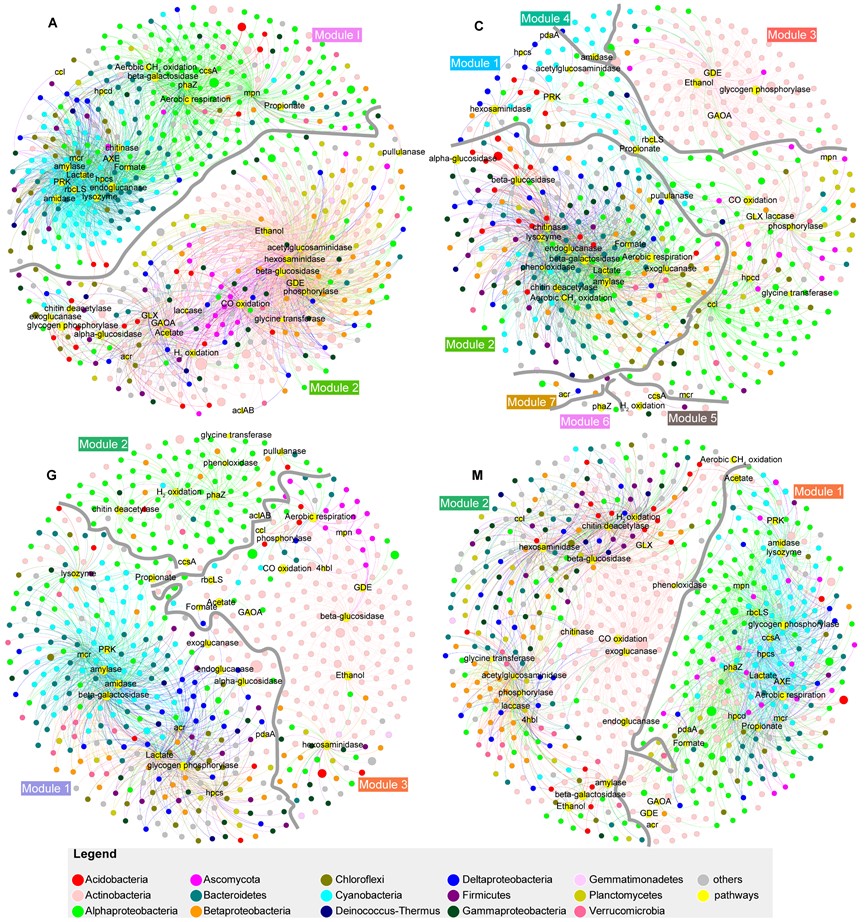
Co-occurrence networks for genes related to carbon cycle and high-abundance genera for each biocrust successional stage. (Image by IHB)
(Editor: MA Yun)