
Newsroom
Significant Progress Revealed in Protist 10,000 Genomes Project
Protists, single-celled eukaryotic organisms encompassing unicellular algae and protozoans, inhabit aquatic environments, contributing significantly to ecological balance, material and energy cycles, environmental health, and disease occurrence in both plants and animals. Functioning as primary producers and oxygen generators, they play crucial roles in the carbon cycle and serve as vital sources of human nutrition, bioenergy, and food for aquatic animals. However, they can also pose challenges, leading to harmful algal blooms and red tides, acting as both pathogens and beneficial partners in symbiotic relationships
The NCBI taxonomy system has documented over 60,000 identified protist species, with an unspecified number yet to be discovered. In December 2019, Chinese scientists, spearheaded by the Institute of Hydrobiology (IHB) at the Chinese Academy of Sciences, launched the Protist 10,000 Genomes Project (P10K). The primary objective of this project is to create a comprehensive genetic resource database for protists, aiming to fill the gap in genetic data within this field.
Recently, a collaborative effort between Prof. MIAO Wei's team at the IHB and Prof. ZHANG Zhang's team from the Beijing Institute of Genomics, CAS (China National Center for Bioinformation) resulted in the release of the initial dataset from the P10K project. This data is now available through the P10K database (https://ngdc.cncb.ac.cn/p10k/), and the accompanying research paper has been published in Nucleic Acids Research.
The inaugural data release from the P10K comprises a comprehensive set of 2,959 protist datasets, featuring 1,601 genomes and 1,358 transcriptomes. This dataset encompasses 75% of protist classes and 45% of protist orders. Among these datasets, 1,858 were integrated from public databases, while the P10K team undertook new sequencing for 1,101 datasets, with a primary focus on ciliates. The resultant newly sequenced data contributed to a substantial 37% expansion in the overall size of the protist dataset. Samples collected from diverse Chinese habitats were subjected to single-cell sequencing methods, particularly for protists challenging to culture in a laboratory, constituting 98% of the newly sequenced data.
To overcome the analytical challenges posed by large-scale single-cell omics data, the P10K team developed a standardized analysis pipeline tailored for single-cell sequencing data of protists. This pipeline encompasses assembly, decontamination, species identification, gene annotation, and evaluation processes. Quality assessments reveal that genomes annotated through this pipeline exhibit a similar proportion of medium and high-quality data compared to those available in public databases.
As an integral component of the 10,000 Protists Genomes Project, the P10K database will propel research on eukaryotic origins, diversity, and microbial interactions. Simultaneously, it will facilitate applications of protist genetic resources in ecological conservation, pollutant degradation, nutrition, health, and disease prevention. Furthermore, the database will support the identification of planktonic organisms based on environmental DNA (eDNA), aiding in aquatic ecological health assessments. Of particular significance, the P10K database has forged a close connection between the National Aquatic Biological Resource Center/National Parasite Resource Center (biological resources) and the National Genomics Data Center (genetic resources). This connection is crucial for promoting information interconnection and data sharing across the National Science & Technology Infrastructure of China.
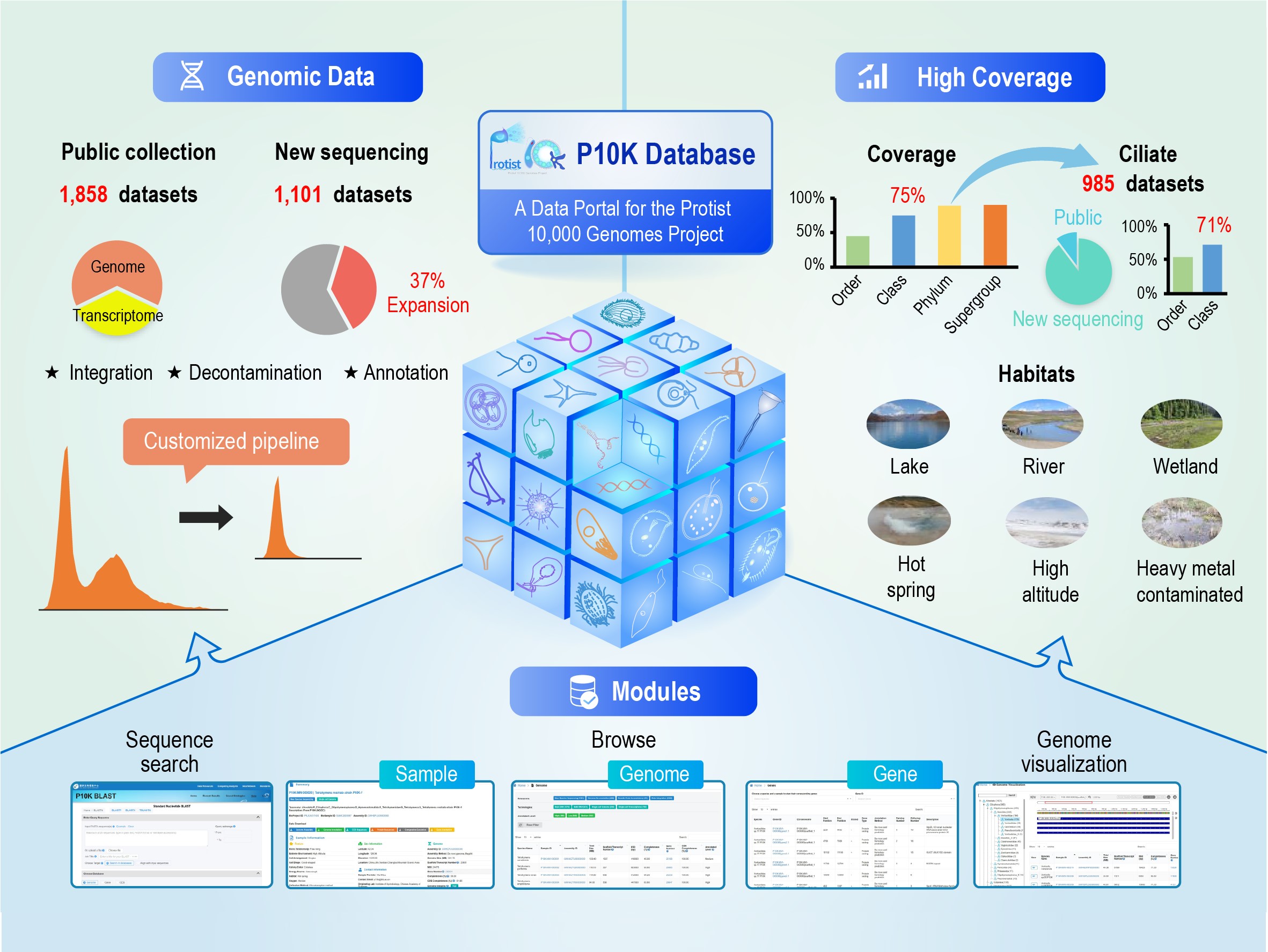
P10K database Contents and Features (Image by IHB)
(Editor: MA Yun)